Plots¶
f_plot¶
Code
1function f_plot(rst,stg,mmf) 2 3% Inform the user that the plots are being done 4disp("Plotting ...") 5 6data_model = mmf.model.data.data_model; 7 8% Import the data on the first run 9load(data_model,'Data','sbtab') 10 11% Generate figure with Scores 12if isfield(rst,'diag') 13 f_plot_scores(rst.diag,stg,sbtab) 14end 15 16% Generate figure with Inputs 17if isfield(rst,'diag') 18 f_plot_inputs(rst.diag,stg,sbtab) 19end 20 21% Generate figure with Outputs 22if isfield(rst,'diag') 23 f_plot_outputs(rst.diag,stg,sbtab,Data,mmf) 24end 25 26% Generate figure with input and Output of all experiments 27if isfield(rst,'diag') 28 f_plot_in_out(rst.diag,stg,sbtab,Data) 29end 30 31% Generate figure with optimization results 32if isfield(rst,'opt') 33 f_plot_opt(rst,stg) 34end 35 36% Generate figures for Sensitivity Analysis 37if isfield(rst,'gsa') 38 f_plot_gsa_sensitivities(rst.gsa,stg,sbtab); 39end 40end
The function that calls all the custom plot functions when appropriate Plots diagnosis that are important to understand if everything is working as it was supposed, it , expected outputs, observed outputs and scores for the models and conditions specified.
Outputs
Figure Scores
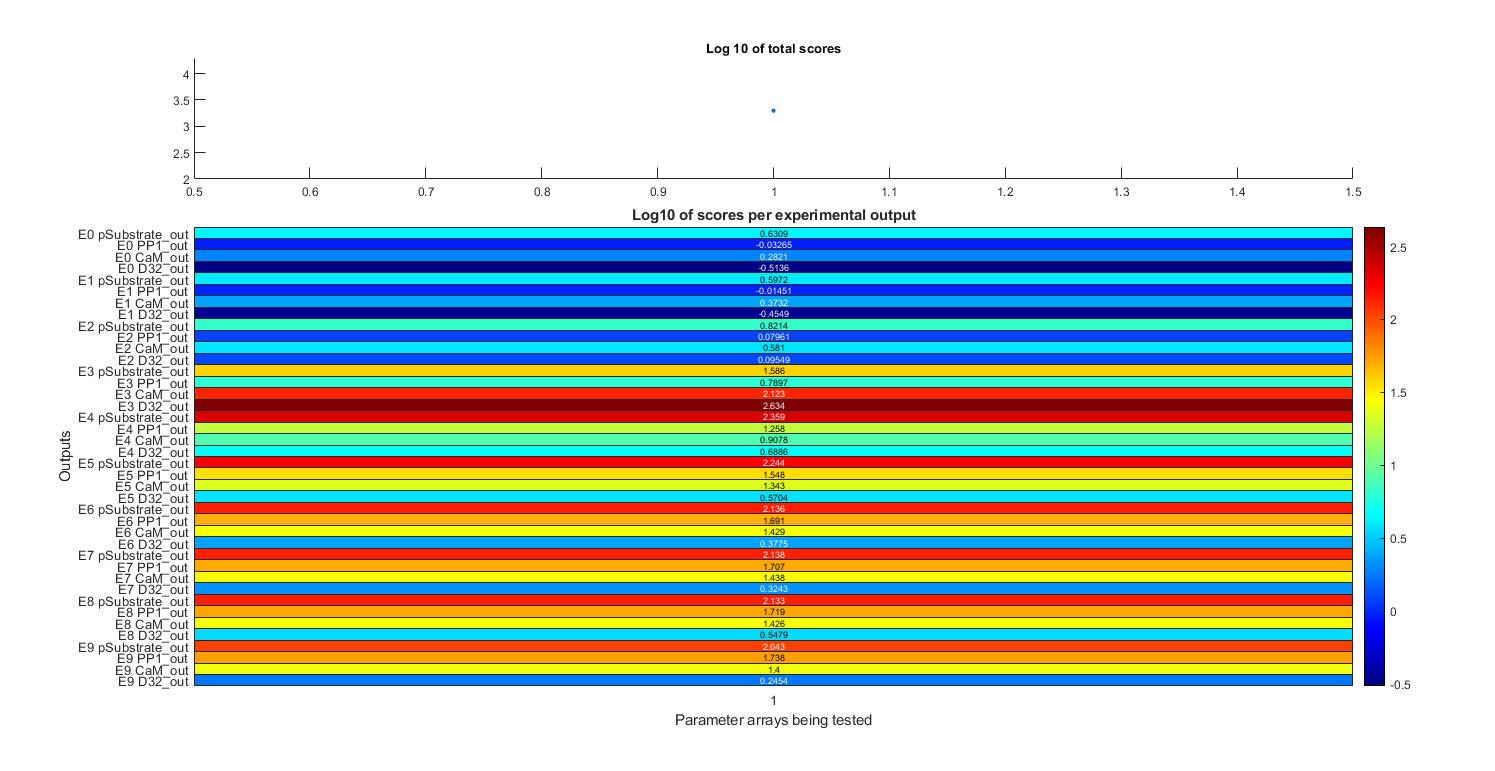
Total scores and scores per dataset given the parameters specified in stg.pa
Code Figure Scores
1function f_plot_scores(rst,stg,sbtab) 2set(0,'defaultTextFontName', 'Helvetica') 3set(0,'defaultAxesFontName', 'Helvetica') 4 5% Generates a figure with Scores 6 7% Inform the user that fig1 is being ploted 8disp("Plotting Scores") 9 10%Closes previous instances of the figure and generates a new one 11figHandles = findobj('type', 'figure', 'name', 'Scores'); 12close(figHandles); 13figure('WindowStyle', 'docked','Name','Scores','NumberTitle', 'off'); 14 15% Generate top plot of figure 1 16subplot(4,1,1) 17 18% Plot the total scores of each parameter array to test 19scatter(stg.pat,[rst(stg.pat).st],20,'filled') 20ylabel('Total Score') 21set(gca,'xtick',[]) 22set(gca,'FontSize',10,'Fontweight','bold') 23 24% Choose correct title according to settings 25if stg.useLog == 1 26 title("Sum of the Log base 10 of the Score of each Experimental Output") 27elseif stg.useLog == 2 28 title("Sum of the Log base 10 of the Score of each Experiment") 29elseif stg.useLog == 3 30 title("Log base 10 of sum of the Score of all Experiments") 31elseif stg.useLog == 4 || stg.useLog == 0 32 title("Sum of the Score of all Experiments") 33end 34 35% Choose the bounds for the x axis so it aligns with the bottom plot 36xlim([min(stg.pat)-0.5 max(stg.pat)+0.5]) 37 38% Generate bottom plot of figure 1 39subplot(4,1,[2,3,4]) 40 41% Generate labels for left of heatmap (Experiment number Dataset number) 42label = []; 43 44% Iterate over the number of experiments 45for n = stg.exprun 46 47 % Iterate over the number of datasets in each experiment 48 for j = 1:size(sbtab.datasets(n).output,2) 49 label{size(label,2)+1} = {strrep("E" + (n-1) + " " + ... 50 string(rst(max(stg.pat)).simd{1,n}.DataNames(... 51 end-size(sbtab.datasets(n).output,2)+j)),"_","\_")}; 52 end 53end 54 55% Choose wether to use the score of each dataset or its log base 10 56% according to settings 57% if stg.useLog == 1 || stg.useLog == 4 58heatline = []; 59 60% Iterate over the number of parameter arrays to test 61for k = stg.pat 62 heatpoint{k} = []; 63 64 % Iterate over the number of experiments 65 for n = stg.exprun 66 67 % Get the score of each dataset 68 heatpoint{k} = [heatpoint{k},rst(k).sd{n,1}]; 69 end 70 71 % Combine heatpoints in order to correctly display heatmap 72 heatline = [heatline;heatpoint{k}]; 73end 74 75% Plot the heatmap 76h = heatmap(transpose(heatline),'Colormap',turbo,'YDisplayLabels',label,... 77 'GridVisible','off','FontSize',10); 78h.CellLabelFormat = '%.2e'; 79 80title("Score of each Experimental Output") 81h.XLabel = 'Parameter arrays being tested'; 82h.YLabel = 'Experimental Outputs'; 83 84end
Figure Inputs

Checks inputs to the model
Code Figure Inputs
1function f_plot_inputs(rst,stg,sbtab) 2% Generates a figure with Inputs, one subplot per experiment 3 4% Inform the user that fig2 is being ploted 5disp("Plotting Inputs") 6 7plot_n = 1; 8fig_n = 0; 9% Iterate over the number of experiments 10for n = stg.exprun 11 12 % Generate the right amount of figures for all plots and calculates 13 % proper subploting position 14 fig_n = f_get_subplot(size(stg.exprun,2),plot_n,fig_n,"Inputs"); 15 16 plot_n = plot_n +1; 17 18 hold on 19 20 % Iterate over the number of inputs in each experiment 21 for j = 1:size(sbtab.datasets(n).input,2) 22 23 % Iterate over the number of parameter arrays to test 24 for m = stg.pat 25 26 % (Until a non broken simulation is found) 27 if rst(m).simd{1,n} ~= 0 28 29 % Plot the inputs to each experiment 30 plot(rst(m).simd{1,n}.Time,rst(m).simd{1,n}.Data(1:end,... 31 str2double(strrep(sbtab.datasets(n).input(j),'S',''))+1),'LineWidth',1.5) 32 33 % Get the correct label for each input of the experiment 34 labelfig2(j) = rst(m).simd{1,n}.DataNames(str2double(... 35 strrep(sbtab.datasets(n).input(j),'S',''))+1); 36 37 ylabel(string(rst(m).simd{1,n}.DataInfo{... 38 str2double(strrep(sbtab.datasets(n).input(j),'S',''))+1,1}.Units)) 39 40 break 41 end 42 end 43 end 44 45 xlabel('seconds') 46 % Add a legend to each plot 47 legend(labelfig2) 48 legend boxoff 49 clear labelfig2 50 51 ylim([0 inf]) 52 53 % Add a title to each plot 54 title("E"+(n-1)) 55 56 hold off 57end 58 59end
Figure Outputs

Expected outputs, observed outputs
Code Figure Outputs
1function f_plot_outputs(rst,stg,sbtab,Data,mmf) 2% Generates a figure with Outputs, one subplot per experimental output 3 4% Inform the user that fig3 is being ploted 5disp("Plotting Outputs") 6 7% Get the total number of outputs to set the total number of plots 8[plot_tn,~] = f_get_outputs(stg,sbtab); 9plot_n = 1; 10fig_n = 0; 11 12% Iterate over the number of experiments 13for n = stg.exprun 14 15 % Iterate over the number of datasets in each experiment 16 for j = 1:size(sbtab.datasets(n).output,2) 17 18 % Generate the right amount of figures for all plots and calculates 19 % proper subploting position 20 fig_n = f_get_subplot(plot_tn,plot_n,fig_n,"Outputs"); 21 22 % Add a legend to the figure 23 if mod(plot_n,24) == 1 24 Lgnd = legend('show'); 25 Lgnd.Position(1) = 0; 26 Lgnd.Position(2) = 0.5; 27 legend boxoff 28 end 29 30 plot_n = plot_n + 1; 31 32 hold on 33 34 % Iterate over the number of parameter arrays to test 35 for m = stg.pat 36 % (Until a non broken simulation is found) 37 if rst(m).simd{1,n} ~= 0 38 39 time = rst(m).simd{1,n}.Time; 40 data = Data(n).Experiment.x(:,j); 41 42 data_SD = Data(n).Experiment.x_SD(:,j); 43 44 % Plot the outputs to each dataset (new subplots) as they 45 % are given in the data provided in sbtab 46 scatter(time,data,'filled','k',... 47 'DisplayName','data') 48 49 errorbar(time,data,data_SD, 'vertical', 'k', 'LineStyle', 'none','LineWidth',1); 50 51 break 52 end 53 end 54 55 % Iterate over the number of parameter arrays to test 56 for m = stg.pat 57 58 % Plot only if the simulation was successful 59 if rst(m).simd{1,n} ~= 0 60 61 time = rst(m).simd{1,n}.Time; 62 [sim_results,~] = f_normalize(rst(m),stg,n,j,mmf); 63 if stg.simdetail 64 time_detailed = rst(m).simd{1,n+2*stg.expn}.Time; 65 [~,sim_results_detailed]= f_normalize(rst(m),stg,n,j,mmf); 66 end 67 68 % Plot the outputs to each dataset (new subplots) and 69 % parameter array to test that are simulated using 70 % Simbiology 71 if stg.simdetail 72 plot(time_detailed,... 73 sim_results_detailed,'DisplayName',... 74 string("Parameter set "+m),'LineWidth',1.5) 75 else 76 77 plot(time,... 78 sim_results,'DisplayName',... 79 string("Parameter set "+m),'LineWidth',1.5) 80 end 81 82 ylabel(string(rst(m).simd{1,n}.DataInfo{end-... 83 size(sbtab.datasets(n).output,2)+j,1}.Units)) 84 end 85 end 86 87 hold off 88 89 xlabel('seconds') 90 91 if stg.simdetail 92 ylim([min([0,min(sim_results_detailed),min(sim_results),min(data-data_SD),min(data)]) inf]) 93 else 94 ylim([min([0,min(sim_results),min(data-data_SD),min(data)]) inf]) 95 end 96 97 % Choose correct title according to settings 98 if stg.plotoln == 1 99 title("E" + (n-1) + " " +... 100 strrep(string(sbtab.datasets(n).output_name{1,j}),'_','\_')) 101 else 102 title("E" + (n-1) + " " +... 103 string(sbtab.datasets(n).output{1,j})) 104 end 105 106 % Choose number of decimal places for y axis 107 ytickformat('%.2g') 108 end 109end 110end
Figure Input and Outputs per experiment

Combined figure of the inputs and outputs for each experiment, on the left side we have the inputs of the experiment and on the right side the outputs
Code Figure Input and Outputs
1function f_plot_in_out(rst,stg,sbtab,Data) 2% Generates a figure with input and Output of all experiments on the left 3% side it plots the inputs of the experiment and on the right side it plots 4% the outputs 5 6for n = stg.exprun 7 8 helper = 1; 9 f_plot_in_out_left(rst,stg,sbtab,helper,... 10 size(sbtab.datasets(n).output,2) > 4) 11 12 for j = 1:size(sbtab.datasets(n).output,2) 13 14 if j/4 > helper 15 helper = helper +1; 16 f_plot_in_out_left(rst,stg,sbtab,helper,... 17 size(sbtab.datasets(n).output,2) > 4) 18 end 19 20 if size(sbtab.datasets(n).output,2) == 1 21 subplot(1,2,j+ceil(j/(2/2))*1) 22 elseif size(sbtab.datasets(n).output,2) == 2 23 subplot(2,2,j+ceil(j/(2/2))*1) 24 elseif size(sbtab.datasets(n).output,2) > 2 &&... 25 size(sbtab.datasets(n).output,2) <= 4 26 subplot(2,4,j+ceil(j/(4/2))*2) 27 elseif size(sbtab.datasets(n).output,2) > 4 28 subplot(2,4,j+ceil(j/(4/2))*2-helper*8+8) 29 end 30 31 hold on 32 33 % Iterate over the number of parameter arrays to test 34 for m = stg.pat 35 % (Until a non broken simulation is found) 36 if rst(m).simd{1,n} ~= 0 37 38 % Plot the outputs to each dataset (new subplots) as they 39 % are given in the data provided in sbtab 40 41 time = rst(m).simd{1,n}.Time; 42 data = Data(n).Experiment.x(:,j); 43 data_SD = Data(n).Experiment.x_SD(:,j); 44 45 scatter(time,data,'filled','k',... 46 'DisplayName','data') 47 48 errorbar(time,data,data_SD, 'vertical', 'k', 'LineStyle', 'none','LineWidth',1); 49 break 50 end 51 end 52 % Iterate over the number of parameter arrays to test 53 for m = stg.pat 54 55 % Plot only if the simulation was successful 56 if rst(m).simd{1,n} ~= 0 57 58 time = rst(m).simd{1,n}.Time; 59 [sim_results] = f_normalize(rst(m),stg,n,j); 60 61 62 if stg.simdetail 63 time_detailed = rst(m).simd{1,n+2*stg.expn}.Time; 64 [~,sim_results_detailed]= f_normalize(rst(m),stg,n,j); 65 end 66 67 % Plot the outputs to each dataset (new subplots) and 68 % parameter array to test that are simulated using 69 % Simbiology 70 if stg.simdetail 71 plot(time_detailed,... 72 sim_results_detailed,'DisplayName',... 73 string("Parameter set "+m),'LineWidth',1.5) 74 else 75 plot(time,... 76 sim_results,... 77 'DisplayName',string("Parameter set "+m),... 78 'LineWidth',1.5) 79 end 80 81 ylabel(string(rst(m).simd{1,n}.DataInfo{end-... 82 size(sbtab.datasets(n).output,2)+j,1}.Units),... 83 'FontSize', 12,'Fontweight','bold') 84 end 85 end 86 87 hold off 88 89 set(gca,'FontSize',12,'Fontweight','bold') 90 91 if stg.simdetail 92 ylim([min([0,min(sim_results_detailed),min(sim_results),min(data-data_SD),min(data)]) inf]) 93 else 94 ylim([min([0,min(sim_results),min(data-data_SD),min(data)]) inf]) 95 end 96 97 xlabel('seconds','FontSize', 12,'Fontweight','bold') 98 99 % Choose correct title according to settings 100 if stg.plotoln == 1 101 title(strrep(string(sbtab.datasets(n).output_name{1,j}),'_',... 102 '\_'),'FontSize', 16,'Fontweight','bold') 103 else 104 title(string(sbtab.datasets(n).output{1,j}),'FontSize', 16,... 105 'Fontweight','bold') 106 end 107 108 % Choose number of decimal places for y axis 109 ytickformat('%.2g') 110 end 111end 112 113 function f_plot_in_out_left(rst,stg,sbtab,helper,reuse) 114 if reuse 115 figHandles = findobj('type', 'figure', 'name', "E " + (n-1) +... 116 " " + helper); 117 close(figHandles); 118 figure('WindowStyle', 'docked','Name', "E " + (n-1)+ " " +... 119 helper,'NumberTitle', 'off'); 120 sgtitle( "Experiment " + (n-1) + " " + helper + " (E " +... 121 (n-1) + " " + helper +")",'FontSize', 28); 122 else 123 figHandles = findobj('type', 'figure', 'name', "E " + (n-1)); 124 close(figHandles); 125 figure('WindowStyle', 'docked','Name', "E " + (n-1),... 126 'NumberTitle', 'off'); 127 sgtitle( "Experiment " + (n-1) + " (E " + (n-1) +... 128 ")",'FontSize', 28); 129 end 130 131 subplot(2,4,[1,2,5,6]) 132 133 hold on 134 for o = 1:size(sbtab.datasets(n).input,2) 135 for p = stg.pat 136 137 % (Until a non broken simulation is found) 138 if rst(p).simd{1,n} ~= 0 139 % Plot the inputs to each experiment 140 plot(rst(p).simd{1,n}.Time,rst(p).simd{1,n}.Data(1:end,... 141 str2double(strrep(sbtab.datasets(n).input(o),'S','')... 142 )+1),'LineWidth',1.5) 143 144 % Get the correct label for each input of the 145 % experiment 146 labelfig2(o) = rst(p).simd{1,n}.DataNames(str2double(... 147 strrep(sbtab.datasets(n).input(o),'S',''))+1); 148 149 ylabel(string(rst(p).simd{1,n}.DataInfo{... 150 str2double(strrep(sbtab.datasets(n).input(o),'S',''))+1,1}.Units),... 151 'FontSize', 12,'Fontweight','bold') 152 153 154 break 155 end 156 end 157 158 end 159 160 set(gca,'FontSize',12,'Fontweight','bold') 161 162 xlabel('seconds','FontSize', 12,'Fontweight','bold') 163 % Add a legend to each plot 164 legend(labelfig2,'FontSize', 16,'Fontweight','bold') 165 legend boxoff 166 clear labelfig2 167 168 ylim([0 inf]) 169 170 % Add a title to each plot 171 title("Inputs",'FontSize', 18,'Fontweight','bold') 172 173 hold off 174 end 175end
Figure Sensitivity Analysis \(S_{i}\)

Figure Sensitivity Analysis \(S_{Ti}\)
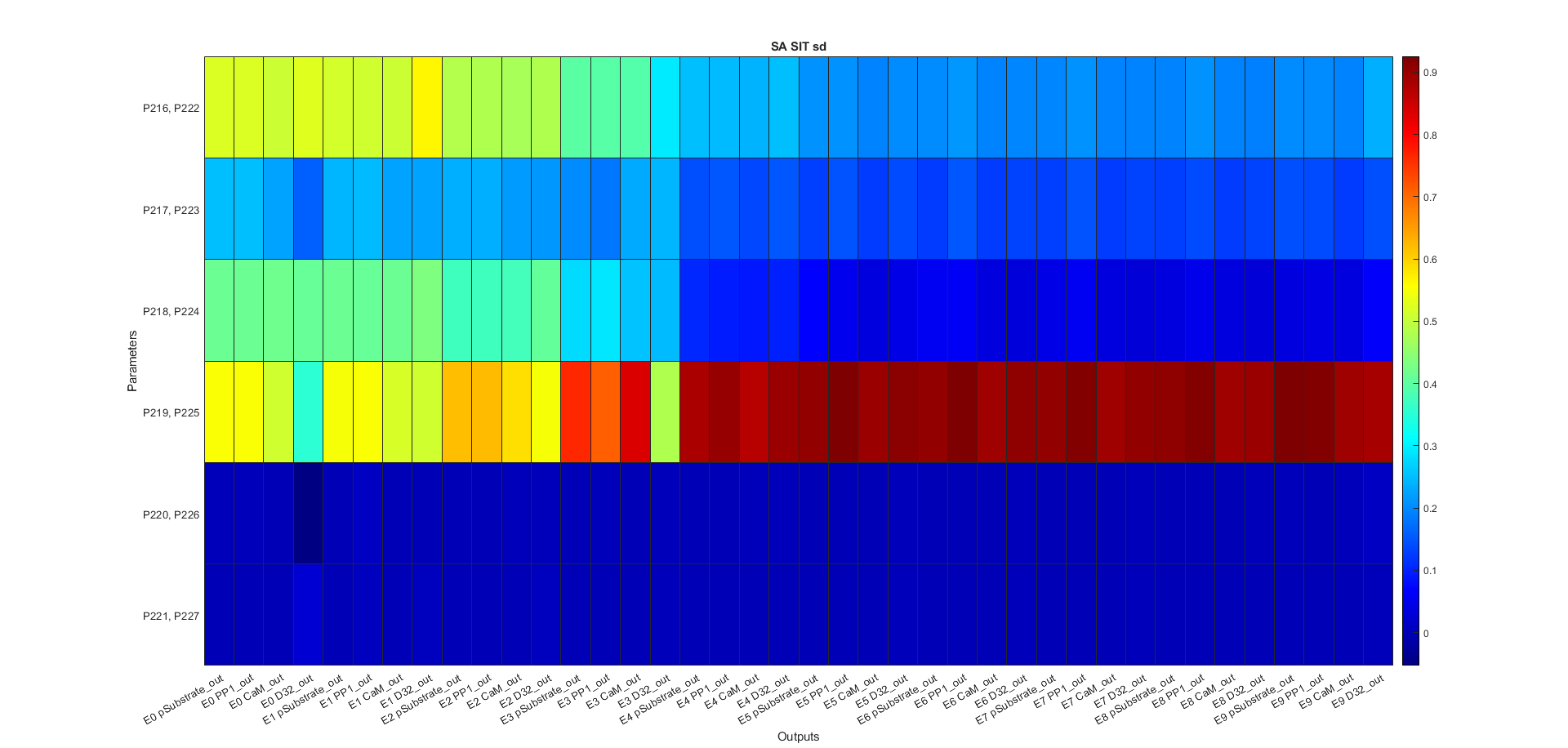
Code figures SA
1function f_plot_gsa_sensitivities(rst,stg,sbtab) 2% Generates figures for Sensitivity Analysis 3 4% Get the total number of outputs 5[~,outputNames.sd] = f_get_outputs(stg,sbtab); 6 7for n = 1:size(outputNames.sd,2) 8 outputNames.sd{n}{:} = strrep(outputNames.sd{n}{:},"_","\_"); 9end 10for n = stg.exprun 11 outputNames.se{n} = "E " + string(n-1); 12end 13 14outputNames.xfinal = outputNames.sd; 15 16parNames = cell(1,stg.parnum); 17parNames2 = cell(1,stg.parnum); 18 19for n = 1:stg.parnum 20 parNames{n} = char("P" + find(stg.partest==n)); 21end 22 23for n = 1:size(parNames,2) 24 parNames2{n} = string(parNames{n}(1,:)); 25 for m = 2:size(parNames{n},1) 26 parNames2{n} = string(parNames2{n}) + ", " +... 27 string(parNames{n}(m,:)); 28 end 29end 30 31% Bootstrapping quartile mean of first order Sensitivity index for score 32% per Experimental Output 33f_generate_plot(rst,stg,outputNames,parNames2,... 34 "Si seo bm",... 35 "First order Sensitivities calculated using the Score of each Experimental Output (Bootstrapping Mean)",... 36 "outputNames.sd",... 37 "transpose(reshape(mean(rst.SiQ.sd(:,:,1:stg.parnum)),[size(rst.SiQ.sd,2),size(rst.SiQ.sd,3)]))") 38 39% Bootstrapping quartile mean of total order Sensitivity index for score 40% per Experimental Output 41f_generate_plot(rst,stg,outputNames,parNames2,"SiT seo bm",... 42 "Total order Sensitivities calculated using the Score of each Experimental Output (Bootstrapping Mean)",... 43 "outputNames.sd",... 44 "transpose(reshape(mean(rst.SiTQ.sd(:,:,1:stg.parnum)),[size(rst.SiQ.sd,2),size(rst.SiQ.sd,3)]))") 45 46% Bootstrapping quartile mean of first order Sensitivity index for score 47% per Experiments 48f_generate_plot(rst,stg,outputNames,parNames2,"Si se bm",... 49 "First order Sensitivities calculated using the Score of each Experiment(Bootstrapping Mean)",... 50 "outputNames.se",... 51 "transpose(reshape(mean(rst.SiQ.se(:,:,1:stg.parnum)),[size(rst.SiQ.se,2),size(rst.SiQ.se,3)]))") 52 53% Bootstrapping quartile mean of total order Sensitivity index for score 54% per Experiments 55f_generate_plot(rst,stg,outputNames,parNames2,"SiT se bm",... 56 "Total order Sensitivities calculated using the Score of each Experiment (Bootstrapping Mean)",... 57 "outputNames.se",... 58 "transpose(reshape(mean(rst.SiTQ.se(:,:,1:stg.parnum)),[size(rst.SiQ.se,2),size(rst.SiQ.se,3)]))") 59 60% Bootstrapping quartile mean of first order Sensitivity index for the 61% final points of the simulations for the output beeing measured 62f_generate_plot(rst,stg,outputNames,parNames2,"Si xfinal bm",... 63 "First order Sensitivities calculated using the final value of each Experimental Output (Bootstrapping Mean)",... 64 "outputNames.xfinal",... 65 "transpose(reshape(mean(rst.SiQ.xfinal(:,:,1:stg.parnum)),[size(rst.SiQ.xfinal,2),size(rst.SiQ.xfinal,3)]))") 66 67% Bootstrapping quartile mean of total order Sensitivity index for the 68% final points of the simulations for the output beeing measured 69f_generate_plot(rst,stg,outputNames,parNames2,"SiT xfinal bm",... 70 "Total order Sensitivities calculated using the final value of each Experimental Output (Bootstrapping Mean)",... 71 "outputNames.xfinal",... 72 "transpose(reshape(mean(rst.SiTQ.xfinal(:,:,1:stg.parnum)),[size(rst.SiQ.xfinal,2),size(rst.SiQ.xfinal,3)]))") 73 74figHandles = findobj('type', 'figure', 'name', 'Si,SiT'); 75close(figHandles); 76figure('WindowStyle', 'docked','Name','Si,SiT', 'NumberTitle', 'off'); 77 78for n = 1:size(parNames2,2) 79 a{n} = char(parNames2{n}); 80end 81 82a = categorical(a,a); 83 84bar(a,[transpose(rst.Si.st(:,1:stg.parnum)),... 85 transpose(rst.SiT.st(:,1:stg.parnum))]) 86xlabel('Parameters'); 87ylabel('Sensitivity'); 88title('Sensitivities calculated using the sum of the Score of all Experiments'); 89legend({'SI','SIT'}); 90legend boxoff 91 92figHandles = findobj('type', 'figure', 'name', 'Si,SiT b'); 93close(figHandles); 94figure('WindowStyle', 'docked','Name','Si,SiT b', 'NumberTitle', 'off'); 95 96T = []; 97 98for n = 1:size(a,2) 99 for m = 1:size(rst.SiQ.st(:,n),1) 100 T = [T;table(rst.SiQ.st(m,n),a(n),"Si")]; 101 end 102end 103for n = 1:size(a,2) 104 for m = 1:size(rst.SiTQ.st(:,n),1) 105 T = [T;table(rst.SiTQ.st(m,n),a(n),"SiT")]; 106 end 107end 108 109boxchart(T.Var2,T.Var1,'GroupByColor',T.Var3,'MarkerStyle','.','JitterOutliers','on') 110xlabel('Parameters'); 111ylabel('Sensitivity'); 112title('Sensitivities calculated using the sum of the Score of all Experiments (Bootstrapping)'); 113legend({'Si','SiT'},'Location','best'); 114legend boxoff 115end 116 117function f_generate_plot(rst,stg,outputNames,parNames2,name,title,... 118 helprer1,helprer2) 119 120eval("figHandles = findobj('type', 'figure', 'name', '" + name + "');") 121close(figHandles); 122eval("figure('WindowStyle', 'docked','Name','" + name +... 123 "','NumberTitle', 'off');") 124 125heatmap_fixer = eval(helprer1); 126heatmap_fixer=heatmap_fixer(~cellfun('isempty',heatmap_fixer)); 127 128heatmap_fixer2 = eval(helprer2); 129heatmap_fixer2 = heatmap_fixer2(:,all(~isnan(heatmap_fixer2))); 130 131h = heatmap(heatmap_fixer,parNames2,heatmap_fixer2,'Colormap',turbo,... 132 'ColorLimits',[0 1],'GridVisible','off'); 133h.CellLabelFormat = '%.2f'; 134 135eval(" h.Title = """ + title + """;") 136h.XLabel = 'Outputs'; 137h.YLabel = 'Parameters'; 138end
Calls
Loads - data.mat
f_get_subplot¶
Code
1function fig_n = f_get_subplot(plot_tn,plot_n,fig_n,fig_name) 2 3size_x = 4; 4size_y = 6; 5size_t = 24; 6ratio_1 = 2; 7ratio_2 = 3; 8 9% If the amount of plots is bigger thatn the maximum amount of plots per 10% figure subdivide the plots to more than one figure 11if plot_tn > 24 12 13 % Generate a new figure for the first plot and each time the number of 14 % plots is greater than figure number divided by max plot number per 15 % figure 16 if mod(plot_n-1,24) == 0 17 18 fig_n = fig_n + 1; 19 20 %Close previous instances of the figure and generates a new one 21 figHandles = findobj('type', 'figure', 'name', fig_name + " " + fig_n); 22 close(figHandles); 23 figure('WindowStyle', 'docked','Name', fig_name + " " + fig_n,'NumberTitle', 'off'); 24 sgtitle(fig_name + " " + fig_n); 25 end 26 27 % Get the correct subploting position for each plot 28 if plot_tn/24 < fig_n 29 subplot(ceil(sqrt((plot_tn-(fig_n-1)*24)/6)*2),ceil(sqrt((plot_tn-(fig_n-1)*24)/6)*3),plot_n-(fig_n-1)*24) 30 else 31 subplot(ceil(sqrt(24/6)*2),ceil(sqrt(24/6)*3),plot_n-(fig_n-1)*24) 32 end 33 34else 35 36 % Generate a new figure for the first plot 37 if mod(plot_n-1,24) == 0 38 39 %Close previous instances of the figure and generates a new one 40 figHandles = findobj('type', 'figure', 'name', fig_name); 41 close(figHandles); 42 figure('WindowStyle', 'docked','Name', fig_name, 'NumberTitle', 'off'); 43 sgtitle(fig_name); 44 45 end 46 47 % Get the correct subploting position for each plot 48 subplot(ceil(sqrt(plot_tn/6)*2),ceil(sqrt(plot_tn/6)*3),plot_n) 49 50end 51end
Inputs
Outputs
Calls
Loads