Welcome to the Subcellular workflow documentation!¶
(Under construction - last updated 2021/07/23)
This workflow has been developed to tackle the challenge of building and analyzing biochemical pathway models, combining pre-existing tools and custom-made software. (Santos et al. 2020) (Preprint)
At the root of our implementation is the SBtab format (Lubitz et al. 2016), a file that can store biochemical models and associated data in an easily readable and expandable way.
We have also developed tools to convert the SBtab format into several formats that can be used in MATLAB®, NEURON, STEPS and COPASI.
Using MATLAB® we have developed custom scripts for parameter estimation, and global sensitivities analysis, as well as diagnostics tools that can be used for model development. The global sensitivity analysis algorithm is modified from Halnes et al 2009.
We demonstrate all these features using three example models, the main one being a modified version of the D1 MSN subcellular cascade model from Nair et al. 2016.
Code and files to run these models in different simulators:
- For Neuron
- For the Subcellular application (STEPS)
- Copasi
Features:
Wrapper for model simulation in MATLAB® (Matlab/Run_main.m)
Analysis of selected parameter sets, using MATLAB® (Matlab/Run_main.m)
Parameter optimization, using MATLAB® (Matlab/Run_main.m)
Global Sensitivity analysis, using MATLAB® (Matlab/Run_main.m)
Conversion tools:
- SBtab (.xlsx,.xls) to SBtab (.tsv), using MATLAB® (Matlab/Run_main.m)
- SBtab (.xlsx) to MATLAB® SimBiology® (.m, .sbproj), using MATLAB® (Matlab/Run_main.m)
- MATLAB® SimBiology® to SBML (.xml), using MATLAB® (Matlab/Run_main.m)Needs to be fixed with our R script (https://github.com/a-kramer/simbiology-sbml-fix)
- SBtab (.tsv) to VFGEN (.vf), using R (https://github.com/a-kramer/SBtabVFGEN)
- SBtab (.tsv) to Mod (.mod), using R (https://github.com/a-kramer/SBtabVFGEN)
- SBtab (.tsv) to SBML (.xml), using R (https://github.com/a-kramer/SBtabVFGEN)
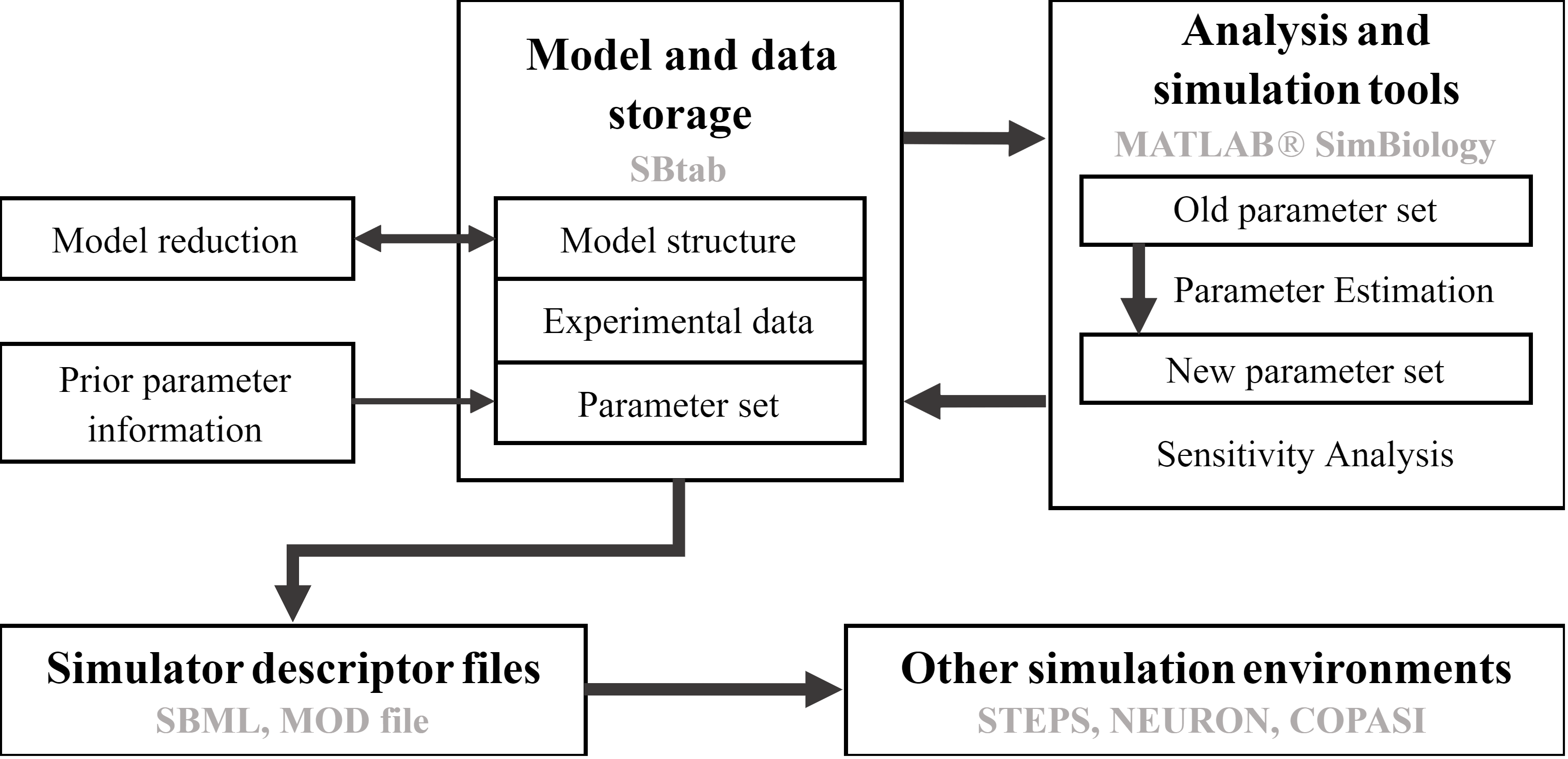
Sections of the workflow in external repositories¶
Conversion tools:
Implemented models¶
Compatibility¶
Subcellular workflow MATLAB® code is compatible with MATLAB® 2020a or above running on Microsoft Windows, macOS and Linux.
Matlab® packages needed:
Optimization Toolbox™
Statistics and Machine Learning Toolbox™
Fuzzy Logic Toolbox™
Financial Toolbox™
Global Optimization Toolbox
SimBiology®
Parallel Computing Toolbox™